Tiny, long-armed dinosaur leads to rethink of dinosaur miniaturization
Small size seems to have come before a change in diet for a tiny dinosaur lineage.
Alvarezsaurids were mostly small-bodied theropods that paleontologists originally misinterpreted as early flightless birds, only to later recognize them as an ant-eating lineage of non-avian dinosaurs. For years, we suspected that Alvarezsaurids underwent a rare process of evolutionary miniaturization directly coupled to a diet of social insects like ants and termites. It was a tidy hypothesis: They got smaller to become more efficient at catching ants.
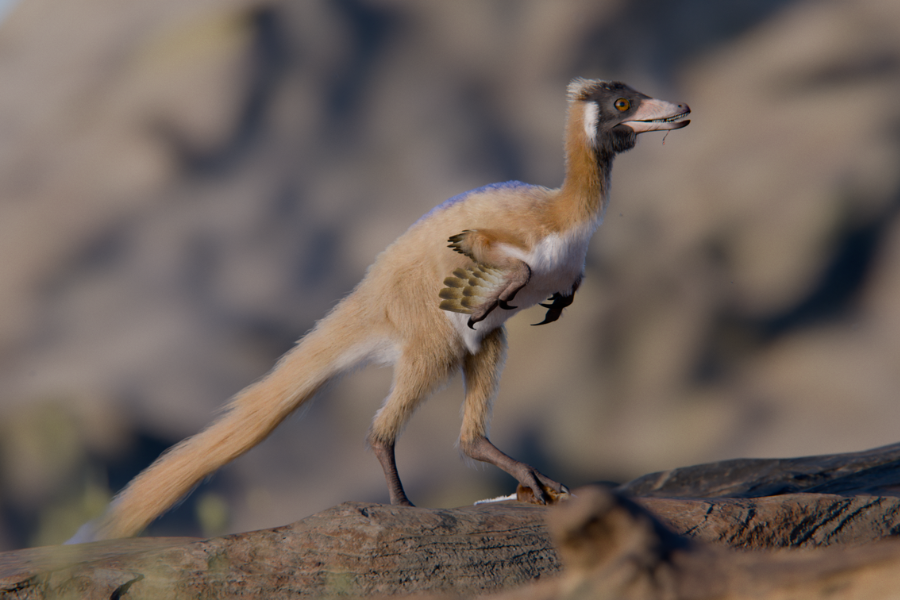
Now, a recently discovered fossil of one of the smallest alvarezsaurids ever found suggests that the evolution of miniature dinosaurs likely wasn’t as neat and linear as we thought. This new species, called Alnashetri cerropoliciensis, probably did not feed on ants at all. “It was a pursuit predator actively hunting insects and small mammals,” said Peter Makovicky, a paleontologist at the University of Minnesota.
The oddball
Alverezsaurids, found mostly in the Late Cretaceous rocks of Asia and South America, had short forelimbs tipped with a single oversized thumb claw built for digging. They also had minute teeth and sensory adaptations akin to those in modern nocturnal birds—everything necessary to work on termite mounds. “The explanation of their small body size has been tied to this specialization,” Makovicky explained.
The dinosaur he and his colleagues found, however, did not look like a specialized ant-eater.
The fossil of Alnashetri cerropoliciensis was unearthed from the Candeleros Formation at the Cerro Policía locality in Argentina’s Río Negro Province and is estimated to have lived roughly 90 million years ago. It currently stands as the most complete and smallest Alvarezsaurid skeleton found in South America.
While missing its skull roof, parts of its right arm, its lower right leg, and much of its tail, the skeleton preserves plenty of its crucial anatomy. Its bone tissue reveals that the alvarezsaurid was a subadult, likely approaching sexual maturity, as indicated by the presence of what appears to be medullary bone, a temporary tissue associated with egg-laying in modern birds. Despite being nearly fully grown, this dinosaur is estimated to have weighed a mere 700 grams.
The real surprise, though, came when researchers realized that Alnashetri wasn’t a highly specialized, late-stage Alvarezsauroid. Instead, despite living in the Late Cretaceous, it occupied an early-branching position among earlier, basal members of the clade.
This combination of tiny size and early-branching status fundamentally breaks our previous model of how these animals evolved. If the miniaturization of Alvarezsauroids was strictly tied to their lifestyle as stubby-armed insect-eaters, an early-diverging species like Alnashetri should have some transitional features on a steady, clade-wide march toward that extreme endpoint. But it didn’t look that way.
“It’s a very long-limbed animal, so it was probably fairly fast. My best analogy would be something like a roadrunner from the American West,” Makovicky said.
Arms and teeth
Late Alvarezsaurids had tiny, robust forelimbs that were less than half the length of their femurs. Alnashetri, though, sported comparatively long forelimbs that were 61 percent of the length of its entire hindlimb. While it had three-fingered hands with a robust first digit, a hallmark of its group, it still retained slender second and third digits, unlike its later cousins.
Other features that challenge the established evolutionary model of miniature dinosaurs are Alnashetri’s jaws and teeth. Its dentition features non-serrated teeth set into sockets, but importantly, these teeth are not extremely small, as they were in the late Alvarezsaurids like Shuvuuia or Jaculinykus. “This decoupled the evolution of small body size from anatomical specializations,” Makovicky explained.
The team concluded that extreme miniaturization in Alvarezsaurids did not necessarily co-evolve with either the evolution of smaller arms more suitable for digging or small teeth built for crushing ants and/or termites. Instead of a clade-wide trend where the entire lineage steadily shrank over time, a new evolutionary model that includes Alnashetri suggests that Alvarezsaurid body mass fluctuated repeatedly. Alnashetri, it turns out, achieved its 700-gram frame independently from the other, highly specialized alvarezsaurid species.
But Alnashetri didn’t just upend the understanding of how Alvarezsaurids evolved their tiny bodies. It also redrew the map of where they lived.
Museum tour
Before Makovicky’s study, it was a mystery why Alvarezsaurids were found almost exclusively in the late Cretaceous rocks of Asia and South America. The previous leading hypothesis suggested that the group must have dispersed back and forth between these two landmasses relatively late in the game. But placing Alnashetri, a remarkably basal member, into their evolutionary tree created a massive ghost lineage. The phylogenetic analysis linked geographically close South American species to much older, geologically distant Asian taxa like Bannykus and Xiyunykus, implying that the group must have diverged way back in the Jurassic period.
To explain this chronological and geographic gap, Makovicky and his colleagues started digging through historical museum collections to see if early Alvarezsaurids had been hiding there under different names. It turned out they had.
The team successfully reidentified a small, fragmentary theropod from the Upper Jurassic Morrison Formation in North America, as well as a Lower Cretaceous taxon from the Isle of Wight in Europe. These were early, diverging Alvarezsaurids, and they possessed distinct features such as specialized ball-and-socket joints in the neck vertebrae that are unique in the Alvarezsaurid clade. These museum reidentifications entirely changed the biogeographical story.
If Alvarezsaurids were roaming North America and Europe in the Jurassic and Early Cretaceous, they weren’t just performing a late-stage migration between Asia and South America. Instead, the new model proposed by Makovicky and his team reconstructs a widespread Pangaean distribution. Early Alvarezsaurids were likely present across the globe before the supercontinent Pangaea fully fractured.
The Late Cretaceous distributions we see in the fossil record today would therefore be the result of populations slowly becoming isolated as the continents drifted apart, combined with regional extinctions that wiped them out in places like North America and Europe. The populations in Asia and South America represent surviving pockets.
Still, Makovicky’s work produced far more questions than answers. If at least some Alvarezsaurids did not evolve their miniature bodies as an adaptation to eating ants, what made them so small?
Messy evolution
“We sort of falsified this nice narrative where Alvarezsaurid body size change was driven by ecology, but unfortunately, we don’t have anything good to replace it,” Makovicky acknowledged.
The classic story of Alvarezsaurids—a lineage steadily shrinking in lockstep as it committed to a life of termite-hunting, finally migrating across the Late Cretaceous globe—was neat and logical, but it’s apparently gone now. “That’s science. Sometimes you can falsify a hypothesis without necessarily finding a better one to support,” Makovicky added. But his team is already busy looking for evidence documenting the new, more complex and messier version of Alvarezsaurid evolutionary history. “We have a couple of angles we’re pursuing,” he said.
The first involves taking a closer look at Alnashetri’s anatomy using CT scans. The goal here is to treat Alnashetri as a starting point to understand the stepwise evolution of its ant-eating, specialized cousins. Most of this meticulous scanning is currently happening in Argentina. The second angle, though, seems way more thrilling. “By pure luck, we found another Alvarezsaur in the same general area,” Makovicky said.
The other Alvarezsaur is bigger than Alnashetri and has slightly shorter forelimbs. “It’s still being prepared, but I think it will sort of give us the next chapter in the story of how Alvarezsaurids evolved,” Makovicky explained. “It’s probably a few years out in the making.”
Makovicky’s work on Alnashetri is published in Nature: https://doi.org/10.1038/s41586-026-10194-3
Tiny, long-armed dinosaur leads to rethink of dinosaur miniaturization Read More »